NHS Wales Informatics Services has been carrying out a technical evaluation into openEHR to test its viability as a repository for structured clinical data. The technology will be rolled out soon to support national projects such as Accelerating Cancer and to provide a shared medications record for NHS Wales.

What is openEHR?
openEHR (pronounced “open-air”) offers a specification for a vendor neutral approach to open standards based clinical models and software. On top of this, we can build digital patient records and applications. And as it is a specification, openEHR based tools and repositories are available from several vendors but importantly, they all promise compatibility with each other’s products. This means data held in one openEHR Clinical Data Repository (CDR) can be surfaced in a variety of places, natively supporting federated approaches for stakeholder organisations.
The specification describes two important elements;
- the clinical models and
- the rules that govern them.
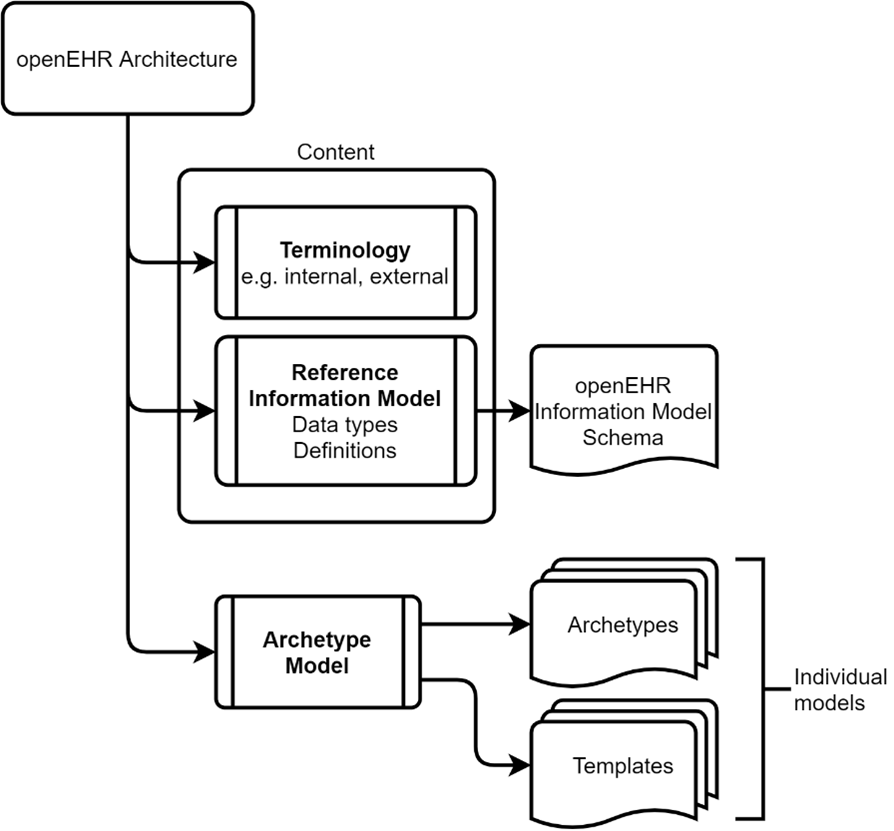
Figure 1 openEHR splits the rules and logic that govern data (i.e. reference information model) from the clinical models themselves.
Traditional approaches to digital record building entwined these two elements and this can cause problems as and when changes are required that affect the database schema. While it is generally agreed that tightly coupling the application to the persistence layer should be avoided, openEHR takes that one step further by dividing the reference model and terminologies from the clinical models (known as archetypes).
The Archetype Model
Archetypes define the possible clinical content, representing a model that originates from actual clinical practice. Archetypes have a governance boundary around them, essentially representing a clinical sign off that can be used to support the idea of a clinical data standard. One or more archetypes can then be used in a template that represents a specific clinical use case.
A simple example could be the development of a Body Mass Index (BMI) app. In order to achieve this, we need three principle archetypes to capture height, weight and the BMI itself. The benefit of using these standard archetypes is that they can also be inserted into other templates that require the same clinical data. And when that data is stored in a common way, it can be queried and reused, reducing the burden of repeated data entry.
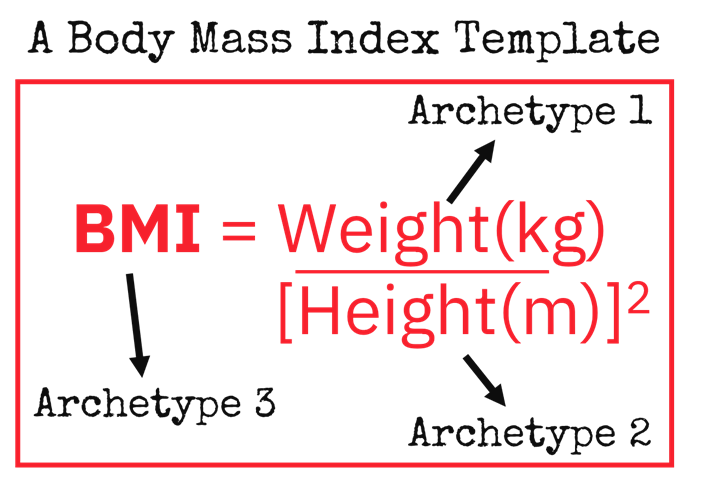
Figure 2 Example use archetypes to form a template.
These models are openly available and have been shown to demonstrate significant reusability for NHS Wales use cases. It has been estimated that approximately 8.5 person years of effort has contributed to the published resources to date (1).
Work with national groups such as Wales Cardiac Network demonstrated effectiveness of the tooling to support the development, publish and review of openEHR models utilising the Clinical Knowledge Manager (CKM) (2). There is however a requirement for clinicians or informaticians that are experienced in clinical care pathways to support curation of new archetypes and this is a challenge to unlock going forward.
Reuse of data
A key attribute of any clinical data repository is its ability to facilitate reuse for additional clinical requirements and for audit and reporting purposes. The Archetype Query Language exists to support this. AQL provides a means of performing queries on the CDR for individual or multiple records at either the patient or archetype level, maintaining data provenance and exporting for more advanced analysis where needed.
Semantic coherence is maintained through the defined reference information model described earlier, with structured data independent of both the application and the underlying database technology that the openEHR resides upon. You can deploy openEHR on a variety of databases from SQL and Postgres to Oracle and MongoDB and AQL will work regardless. An additional benefit is that AQL can be used with the standard RESTful APIs to permit structured compositions or archetype queries to be retrieved in commonly understood formats such as JSON and XML by developers and analysts.
Does it compete with HL7 FHIR?
No, emphatically not but this is a common misconception as both specifications have a certain amount of commonality. But the best way to view the differences is that FHIR represents a standards based approach to messaging (commonly known as technical interoperability) while openEHR supports persistence.
Various elements within the openEHR reference model can tie both together and enable semantic interoperability for clinical applications as it has its own set of APIs. However, HL7 FHIR is recognised as the emerging messaging syntax and the standard for how health organisations and systems can communicate. To do this effectively, FHIR represents a series of generic resources to act as an envelope for a variety of clinical data items. Conversely, openEHR represents the detailed, granular view of that persisted, accessible data.
Why not just send openEHR archetypes between systems?
The answer is complexity. Archetypes and templates represent a given clinical use case and an archetype may be represented in several different templates with varying degrees of completeness.
If we consider recording Blood Pressure, there is a clear difference between the data requirements in primary care versus that of an intensive care ward. The more rudimentary systolic / diastolic observation is appropriate for a GP and therefore these are the two principle data items captured for this scenario. A nurse on a ward may also wish to capture the ‘Cuff size’ to more rapidly facilitate blood pressure readings on a busy ward and this introduces an additional data item. Conversely, an ICU ward may be streaming data directly from a machine capturing the mean arterial pressure and to substantiate a semantic context, the corresponding formula used for the calculation and device serial numbers may also be recorded. All three examples are valid, and they also all conform to a single clinical model.
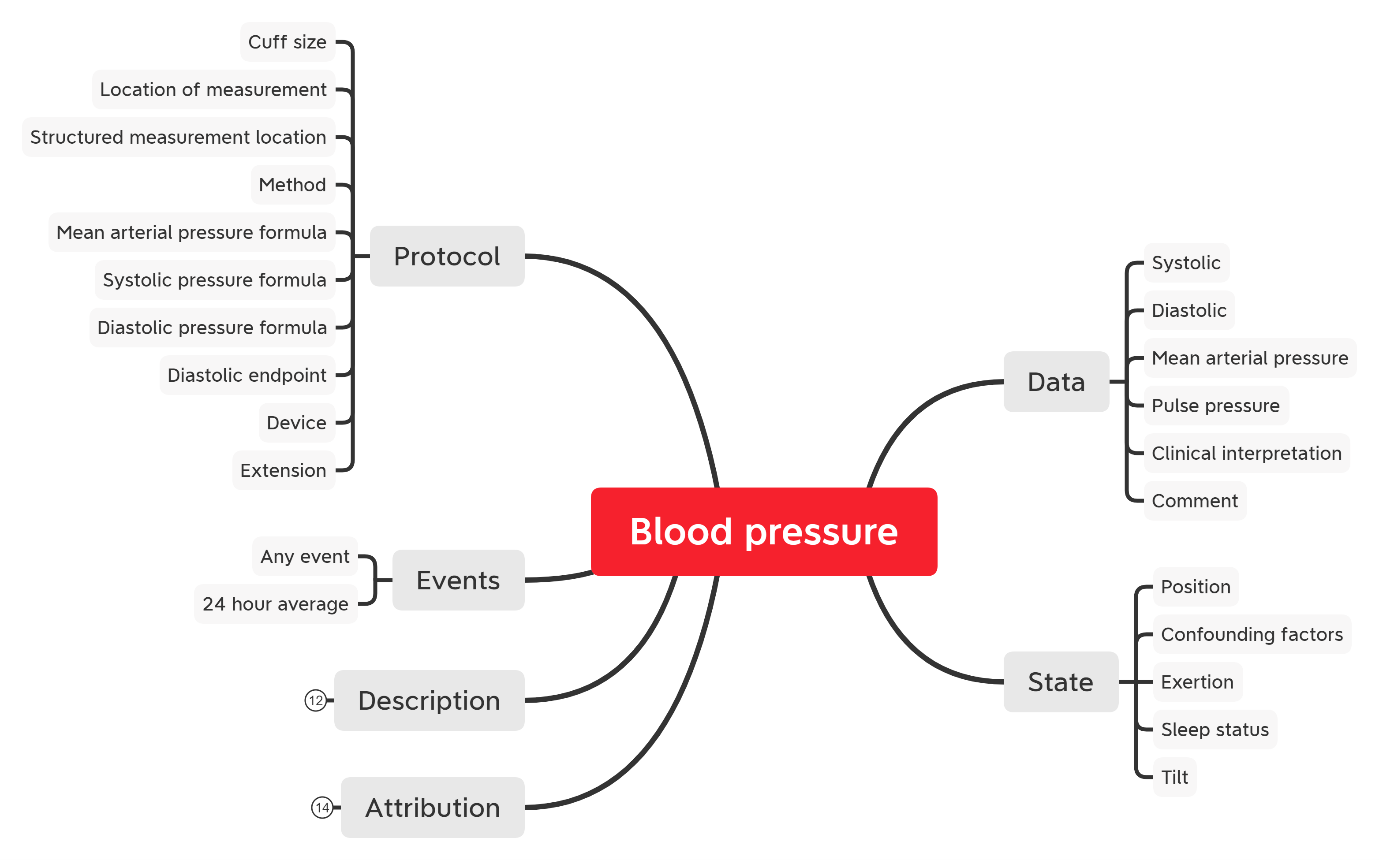
So “completeness” is arbitrary in this regard as system builders are not required to use 100% of the content that may be captured using an archetype. The maximal approach to modelling is designed to represent a holistic view on a given data item, essentially what is feasible to be captured. But that does not mean this has to be done in all scenarios.
But if we wanted to facilitate interoperability with a maximal approach, we would need to ensure that there was either a single all-encompassing FHIR model available that was accepted and understood by all parties, or several APIs that represented each use case. Each would require maintenance and potential development resources whenever a change occurred to that model. Although this is a moot point when many data items may not need to be sent between all systems in the first place.
The requirements for messaging between systems and the underlying persistence of clinical data are not always the same, but both technologies complement each other. It is this “toolbox” approach that we are seeking to employ; utilising the FHIR standard to ensure vendors and organisations are talking in the same language and able to represent the clinical content in a variety of flexible ways. At the same time, that data is persisted according to one agreed model which is the principle function of openEHR.
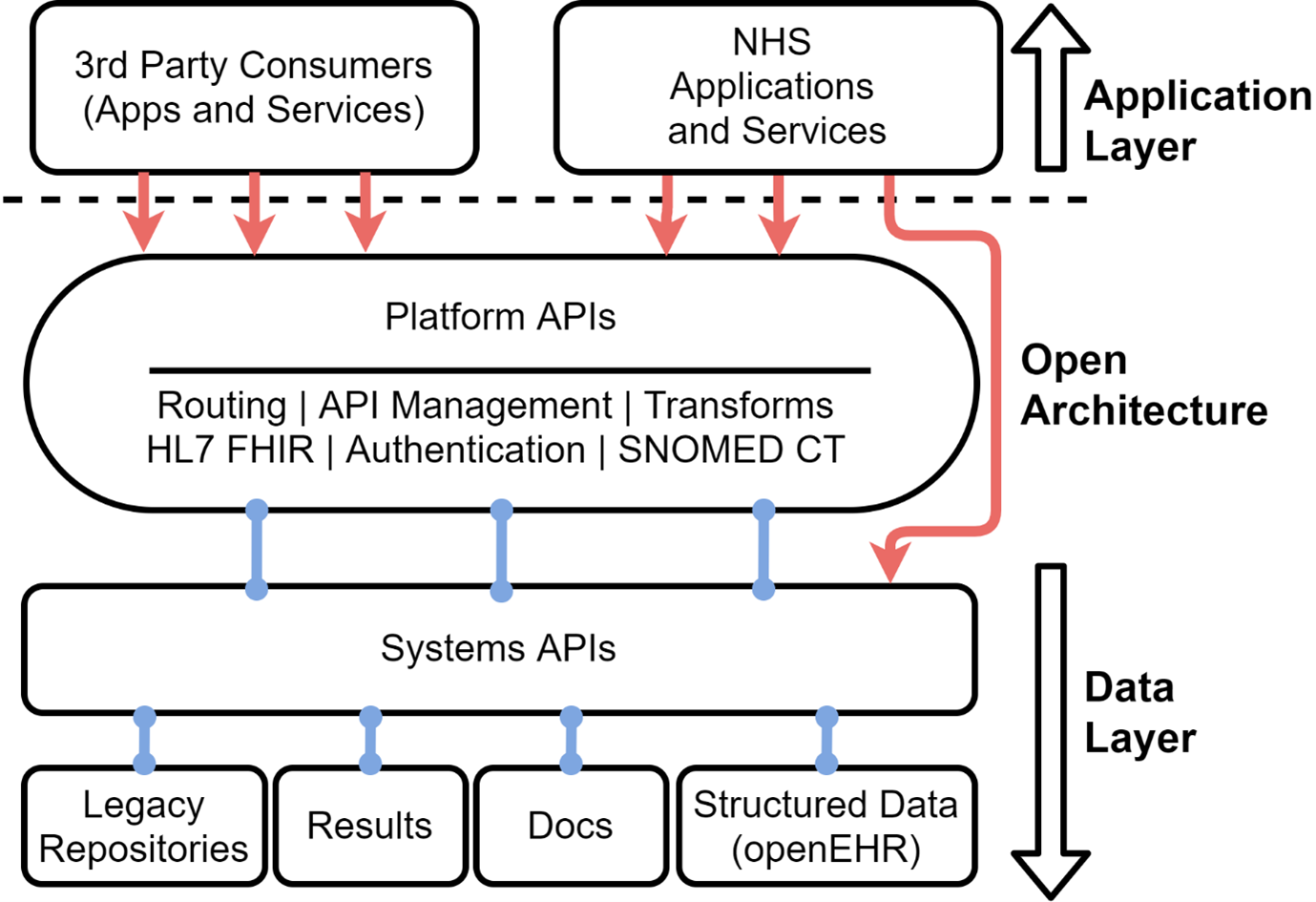
Adopting openEHR allows NHS Wales to start the transition from the siloed and document bound approach to the emerging open platform (3). We can start to with a future first mentality which does not imply complexity. Therefore, we are looking to first deliver a specific use case: the treatment repository to support cancer management requirements.
The above diagram describes how data is stored separately from the applications that collect, edit and display it. The data layer helps deliver content that is standardised in terms of format, nomenclature, terminology, and definition. This in turn allows it to flow into other systems, as specified by the data owner. Good rules for data storage will not change much, if at all, allowing for persistence and, ultimately, increasing operability over time.
The application layer requires a fully systemic design of workflow. This means knowing context, or what comes before and after, in the care journey. It is based on triggered events of care or intervention (e.g., clinical workflows or health consumer alerts) rather than constant human monitoring. This workflow is heavily dependent on the underlying, standardised data structures as the foundational models provided by archetypes provide a baseline from which to build.
The wide-angle view on the adoption of the Open Platform is important, of which openEHR is a component part. It can leverage standard APIs and work with other platform elements (e.g. HL7 FHIR based applications) to form an interoperable façade over a persistent structured data layer. Principles that underpin open platforms (such as reducing vendor lock-in, technological neutrality and open interfaces that support data access) are vital to realising the potential of the National Data Resource and the health data for Citizens of Wales.
Summary
Having given an overview of openEHR, we have just scratched the surface of how it could support present and emerging use cases. Over the next two years we will be collaborating with colleagues in the National Data Resource to expand on these requirements, and working with the All Wales Eye Care Digitisation Programme as they transition Open Eyes to openEHR. And we will be sure to feed back our activity at future DHEW events as we hit each milestone.
Interested in learning more? We're running two openEHR events in February:
- openEHR for NHS Wales, 16th February - an opportunity for attendees to learn about OpenEHR and broaden their understanding of this technology which is being used across a range of organisations in the UK.
- openEHR: a technical overview, 23rd February - an opportunity for attendees to learn about the technology and broaden their understanding, focussing on the technical aspects of its architecture.
- H. Leslie, “State of the clinical modelling program and international CKM,” Atomica Informatics, 2019
- Clinical Knowledge Manager
- Defining an Open Platform, Apperta Foundation
